Bacterial Resistance to Antibiotics
Bacterial resistance to antibiotics
🪫 “Bacterial Resistance to Antibiotics”
One of the most common arguments used by laughing Darwinists is the ability of bacteria to develop resistance to antibiotics,
and they consider this a clear evidence of microevolution …
🔍 Introduction:
Let us first clarify that the main reasons for this resistance are:
-
The transfer of antibiotic resistance genes from bacteria that are already resistant to antibiotics
-
The misuse of antibiotics
-
Or through destructive mutations
But all of this has nothing to do with Darwinian theories .
We will explain how bacteria develop antibiotic resistance in two ways:
✅ Resistance is acquired through the transfer of resistant bacterial genes to another cell that has been decomposed (transformation ).
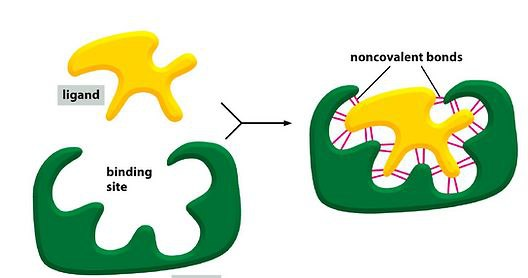
Resistance genes are found on the plasmid,
and these genes produce an enzyme that destroys or inactivates the bacterial-killing substance in the antibiotic,
or they produce selective cellular pumps whose job is to expel toxins, such as the antibiotic.
These genes contain a precise mechanism for integrating them into the other gene,
and this is also against Darwinian randomness.
✅ Through mutations;
But bacteria do not acquire resistance from random mutations,
or from evolutionary mutations that lead to new additions.
As Science Direct mentioned, bacterial transformation (gene transfer) is a programmed process.
(Meaning: Against chance and randomness)
📌 Bacterial transformation is a programmed process resulting in genetic transfer and diversity.
It relies on the development of competence via regulatory circuits which are diverse and tailored to the particular lifestyle of each species.
Despite this diversity, some species have been reported to trigger competence in response to antibiotics.
In another article, I mentioned that it is stably integrated into the recipient cell (also against chance).
Bacterial Transformation , as mentioned above, means the uptake of DNA molecules through the cell wall from the external surroundings, followed by stable incorporation into the recipient genome, or replication as an independent plasmid.
📎 https://www.sciencedirect.com/topics/medicine-and-dentistry/bacterium-transformation
🧬 Let’s start first:
★ Methods of transfer of the resistance gene:
- After the death of the bacterial cell resistant to the antibiotic, its genome remains,
so the living bacterial cell that is not resistant to the antibiotic picks up the gene from the dead cell,
then it is transferred to the living bacteria,
and it is integrated precisely as we explained above —
and this phenomenon is called bacterial transformation .
- By transfer or transduction ,
in which bacterial viruses (bacteriophages ) transfer the resistance gene from the bacteria that possess it to the cell that does not have it.
- Through conjugation ,
where genes are transferred from a donor cell to a recipient cell via small pili that bacteria have — specifically sex pili —
and the resistance gene is transferred through them, and then integrated, of course.
🧬 Integrative and Conjugative Elements (ICEs) play a well-established role in disseminating the genetic information underlying adaptive traits.
- ICEs are mobile DNA (~20 Kbp to >500 Kbp in size )
that contain the genes required for genomic integration, excision, and transfer via conjugation.
In addition, they contain a wide range of gene cargoes conferring phenotypes such as:
Antibiotic resistance
Heavy metal resistance
Nutrient utilization
Pathogenicity
(Reviewed by refs. 8 and 9)
📎 https://www.pnas.org/doi/10.1073/pnas.2214521120
🦠 Now: How do these genes contribute to antibiotic resistance?
These genes disable the antibiotic by producing enzymes that break it down,
such as beta lactamase , which destroys fungal toxins.
They also enable bacteria to reproduce what was targeted by the antibiotic,
such as a vitamin or organic compound important for life and growth.
They also pump harmful compounds out of the cell
as part of the exocytosis system inherent in the bacteria’s creation.
Active efflux from procaryotic as well as eucaryotic cells strongly modulates the activity of a large number of antibiotics. Effective antibiotic transport has now been observed for many classes of drug efflux pumps. Thus, within the group of primary active transporters, predominant in eucaryotes, six families belonging to the ATP-binding cassette superfamily, including the P-glycoprotein in the MDR (Multi Drug Resistance) group and the MRP (Multidrug Resistance Protein), have been recognized as being responsible for antibiotic efflux.
https://www.sciencedirect.com/science/article/abs/pii/S0006295200002914
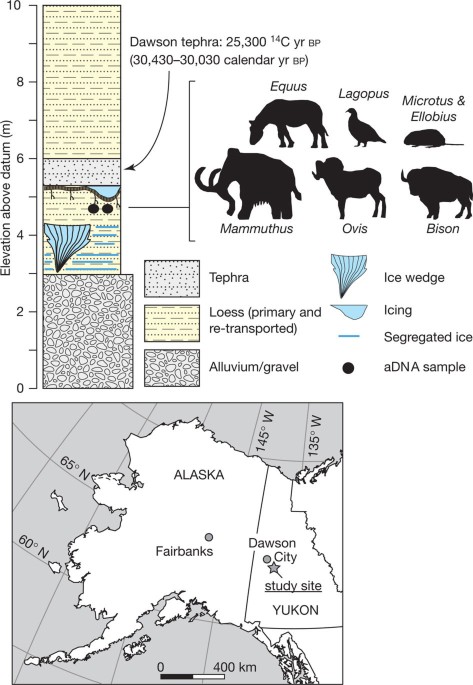
★ Resistance genes use three mechanisms to counteract antibiotics: modification, sequestration, and destruction. ★ Where is the evidence that bacteria are resistant to begin with? There is a 1988 experiment in culturing bacteria found on the frozen bodies of Arctic explorers dating back to 1845. When they awakened these bacteria, they were able to resist antibiotics, which hadn’t even been created yet. In fact, the difference is that antibiotics were invented before they existed, which means they already had this resistance before antibiotics existed. Nature reported that bacteria have been resistant to antibiotics for about 30,000 years. Here we report targeted metagenomic analyses of rigorously authenticated ancient DNA from 30,000-year-old Beringian permafrost sediments and the identification of a highly diverse collection of genes encoding resistance to β-lactam, tetracycline, and glycopeptide antibiotics.
https://www.nature.com/articles/nature10388
By the way, I mentioned that this means they weren’t exposed to selective pressure (natural selection). 😁
These results conclusively show that antibiotic resistance is a natural phenomenon that predates the modern selective pressure of clinical antibiotic use. Another study reported that bacteria have been resistant to antibiotics for at least 5,000 years. Using functional metagenomics to study the resistomes of bacterial communities isolated from different layers of the Canadian high Arctic permafrost, we show that microbial communities harbored diverse resistance mechanisms at least 5,000 years ago. Another site reported that they have been resistant to antibiotics for more than eight centuries since their invention
https://www.sciencedaily.com/releases/2014/02/140223131629.htm
So far, the myth that bacteria can add entirely new mechanisms due to selective pressure has been dispelled.
{Embed}
https://www.nature.com/articles/nature10388
Antibiotic resistance is ancient
Nature - Antibiotic resistance is thought to have evolved long before naturally occurring antibiotics and their derivatives were used to treat human disease, but direct evidence for genes that…
{Embed}
https://www.sciencedaily.com/releases/2014/02/140223131629.htm
‘Microbial Pompeii:’ 1,000-year-old plaque preserves bacteria, micr…
A ‘microbial Pompeii’ has been discovered, preserved on the teeth of skeletons around 1,000 years old. The research team discovered that the ancient human oral cavity carries numerous opportunistic pathogens and that periodontal disease is caused by the same bacteria today as in the past, despite major changes in human diet and hygiene. “T…

🪫 Bacterial Resistance to Antibiotics: Part Two
We’ve previously discussed the existence of mechanisms for bacterial resistance to antibiotics,
and explained that bacteria were either pre-equipped to respond to antibiotics before they were invented,
or they acquired the resistance gene from another bacteria or virus,
or through mutations.
Now, let’s talk about bacterial resistance to antibiotics through mutations .
❓ Is this really evidence of evolution, as evolutionists claim?
When discussing bacterial acquisition of resistance through mutations,
the general public might immediately think that these are mutations that add information to the genetic code,
leading to the addition of defensive and immune mechanisms to the bacteria.
🪫 However, all observed evidence indicates that they are mutations that lead to:
Deterioration
Modification
Destruction
Loss of function
🧪 Let’s explain this further to get the idea across:
A scientist decided to study the ability of the enzyme Beta-Lactamase to bind to a type of antibiotic
through several mutations and intense selective pressure.
🪫 The result was a complete failure.
📎 The results predict, with >99.9% confidence, that even under intense selection the IMP-1 β-lactamase will not evolve to confer increased resistance to imipenem.
Some antibiotics work by binding to certain sites in bacterial cells, called receptor sites ,
or by relying on an enzyme produced by the bacteria that the antibiotic targets.
If a mutation destroys this binding site, the bacteria escapes the antibiotic —
because the receptor it acts on is destroyed.
{Embed}
https://pmc.ncbi.nlm.nih.gov/articles/PMC353146/
In Vitro Evolution Predicts that the IMP-1 Metallo-β-Lactamase Doe…
In vitro evolution was used to predict whether the IMP-1 metallo-β-lactamase has the potential to evolve an increased ability to confer resistance to imipenem. Screening of eight libraries containing 9.8 × 106 ± 1.4 × 106 (mean ± standard error) …

🔌 For example :
Look at the first picture below: What happens if a strong blow hits one of the cavities?
An antibiotic will not be able to bind to the receptor.
Just like an electrical plug that is bent after being hit hard will not fit properly —
What happens? It will not be able to enter its original location, and therefore will not work.
Did we add anything?
❌ No, we have destroyed something that existed in order for it to survive.
However, this mutation simultaneously weakens the function of the enzyme or the binding site,
making the organism weaker and less efficient.
🪫 This is called the “fitness cost.”
This cost is usually high, making the strain that has undergone destructive mutations unable to survive in a natural environment
compared to its competing peers from the original strain that did not have impaired functions.
These destructive mutations may even slow the reproduction rate of the original strain.
💉 Therefore:
One treatment method is to recommend stopping the use of antibiotics for a certain period
if the bacteria develop resistance to them —
this is to restore the original strain that is not resistant to the antibiotic,
and then resume the antibiotic use against the original strain later.
🧬 According to evolution:
Natural selection works to eliminate the individual unable to survive,
because it selects only the fittest.
🪫 And here, the original strain that did not undergo mutations is the fittest to survive.
📎 https://pubmed.ncbi.nlm.nih.gov/18603474/
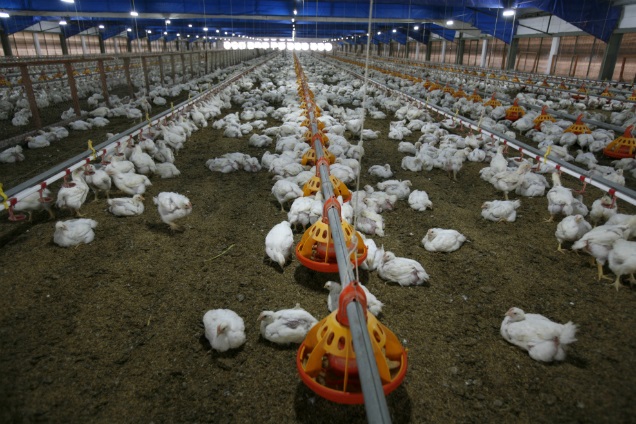
{Embed}
Stop using antibiotics in healthy animals to preserve their effecti…
WHO is recommending that farmers and the food industry stop using antibiotics routinely to promote growth and prevent disease in healthy animals.

{Embed}
https://pubmed.ncbi.nlm.nih.gov/18603474/
Fitness costs of resistance to antimalarial drugs - PubMed
It has been recently reported that the prevalence of mutations associated with chloroquine resistance declined during the dry season. Fitness costs of drug resistance were suggested to be responsible for reduced survival of mutant parasites, and only parasites surviving chronic infections were trans …

Neither is this evolution, nor will natural selection by evolution preserve it for the sake of evolution.
🧠 Another type of mutation is regulatory mutation :
Some antibiotics target essential proteins produced by bacteria.
If a mutation disrupts the regulation of protein production and causes increased production,
bacteria can bypass the effect of the antibiotic on their proteins.
However, the cost of adaptation comes again:
increasing protein production above the normal rate increases the burden on bacteria,
increasing resource consumption…
🪫 And all of this does not add information to the genome.
Mutation of the staphylococcal accessory regulator (sarA) or sigB was found to significantly increase susceptibilities to both antibiotics and in both strains —
in a manner that could not be explained by changes in the MICs.
The impact of a mutation in sarA was comparable to that of a mutation in sigB ,
and greater than the impact observed with any other mutant.
These results suggest that therapeutic strategies targeting sarA and/or sigB
have the greatest potential to facilitate the ability to overcome the intrinsic antibiotic resistance
that defines S. aureus biofilm-associated infections.
📎 https://journals.asm.org/doi/10.1128/aac.02750-15
Mutations also cause antibiotic resistance by affecting components of the cell surface and membrane, such as the transmembrane transporters responsible for the entry of molecules into the cell. This reduces their effectiveness, which in turn reduces the amount of antibiotics entering the cell. However, they also cause deterioration, because just as they reduce the entry of antibiotic molecules, they also reduce the entry of food molecules and other substances used by the cell. This makes them unable to survive against other bacteria, which prevents natural selection from preserving them according to evolution. There are also other mutations that distort the cell wall, preventing the antibiotic from binding to it. However, this distortion also impedes bacterial growth and makes them vulnerable to other bacteria, which prevents natural selection from preserving them according to evolution. For such reasons, this resistance disappears once the antibiotic is discontinued, and the original strain subsequently prevails because the other strain is deteriorated, not evolved. In the end, do you think that all these mutations occur randomly, or as Darwinian evolution claims?! The so-called bacterial SOS system theory has been established, a process by which bacteria “under high environmental stress, such as DNA damage,” summon a highly mutagenic backup DNA polymerase, which increases the mutation rate in the affected region, helping to cope with the problem. Even destructive mutations are not random; they occur in a precise order. The bacterial SOS response kicks in when bacteria experience DNA damage, and helps the organisms correct and survive DNA damage events. This primer provides a foundation for understanding these events.
https://pmc.ncbi.nlm.nih.gov/articles/PMC1174825/
{Embed}
https://pmc.ncbi.nlm.nih.gov/articles/PMC1174825/
After 30 Years of Study, the Bacterial SOS Response Still Surprises Us
The bacterial SOS response kicks in when bacteria experience DNA damage, and helps the organisms correct and survive DNA damage events. This primer provides a foundation for understanding these events.